One of the things about starting your new lab that can keep you up at night, is that suddenly you’re the one with the final say on a lot of things. This can be intimidating when it comes to things like Health & Safety, Ethics, and spending money. But it’s pretty exciting when it comes to publications. You now get to choose how to present yourself to the world, and in this post, I’m going to encourage you to choose a path that is open, collaborative, and will ultimately help others to interpret your work more clearly. A quick warning for those of you on audio – the post works best with pictures.
At the beginning of my career, I knew nothing about data analysis. I’ll freely admit I analysed everything I did using the geometric mean in the first year of my PhD, rather than the more usual arithmetic mean, not realizing it was a completely different thing. At some point somebody mentioned something, I did some googling, and thank goodness changing the mean calculation didn’t change the overall outcome of my experiments. I’d sat in Medical Statistics sessions, and done a couple of classes on stats that somehow always focused on power calculations, but if I’m brutally honest, it made no sense to me. Then my now husband came along, an astrophysicist turned computational biologist, and in the early days of our relationship, I finally admitted I had no idea what I was doing. His explanation of what a t-test was blew my mind, both because it was insanely simple, and because I couldn’t believe no one had ever presented it to me this way before.
His explanation was something like this (it was far too long ago!);
“Pretty much everything in biology is a distribution. You have one group of people, let’s call them 1 and their values, let’s say their height, look like this as a distribution. Then you have another group of people, let’s call them 2, and you think they are different to 1 in height. So you plot their distribution. And all the t-test does, is say are these two distributions separated, or are they on top of each other? And how confident of our judgement can we be?”
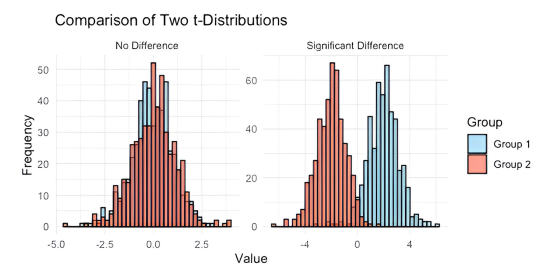
Figure 1 – What’s a t test?
You’re probably giggling at this point about how that was a revelation to me. It is pretty darn simple. But for all those years of staring into space as people re-derived power calculations, no one had ever just used a couple of quick plots to explain this to me. From here, the world opened up.
In general, your statistical test takes a measure of where the middle (usually the busiest) part of your distribution is, followed by a measure of how wide it is (how variable your measures are), and works out how likely it is that things don’t overlap. It does get a bit more complicated, as there are distributions of different shapes, and when there are significant tails to the distribution, especially if the tail is much longer on one side than the other (“skewed”) data. Different distributions are different shapes, so you might choose a different test for that shape, or you might choose to transform your data first to change the shape, to make a test more valid.
Hang on a minute, you say! I don’t come to this blog for stats primers! I come for advice on how to start my own lab! And this is where the two things come together. You have a choice when you write your papers to show that you can do this stuff well, and to really demonstrate to people that they should trust your data. Particularly in the biomarkers field, it is so common for the sole data shown in the paper to be a table of model co-efficients and variables, quite frequently showing only the things the paper says are significant. But how do I know you’ve run a decent model? Did you just not detect my protein, or was it not significant? Without a bit more information, it’s impossible for me to tell if you’ve just cycled through various tests till you found a p value that fits, or if you carefully selected an approach to fit your data. Take this extreme example;
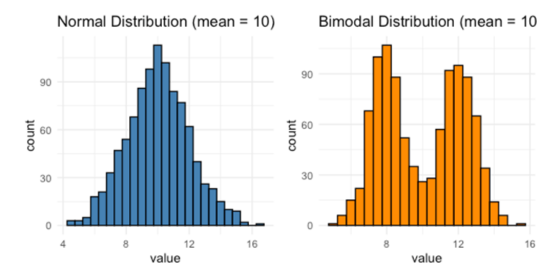
Figure 2 – two very different distributions with the same mean
These two distributions are clearly very different to each other, but they have the same mean value. If you’ve done a test which relies on a normal distribution, such as the plot on the left, it’s probably going to tell you that there’s no difference between the two. And you can see clearly that’s not the case. It therefore really helps me understand your data if the actual datapoints are shown.
See the next example below. On Plot 1, we have some error bars, which suggest that there’s trend towards a difference between the two conditions. In plot 2, we have the data points driving that difference, and it’s super clear that you have a small number of outliers responding to your drug, but that most data points are no different to control. Compare this to plot 3, where the whole distribution is shifted. You see a small difference in error bars, but I’d argue that really undersells the difference between these two distributions. And what experiment you might do next depends on this distribution – either you need to figure out what’s different about your responders, or you may simply need to get a few more datapoints to get a significant effect.

Figure 3 – population shift, or driven by outliers?
So, you’re the boss. You can make sure that you present your data clearly, with all data points shown, and to some extent this allows your readers to do the analysis for themselves. But what about when you’re doing an ‘omics experiment, measuring thousands of things at the same time? You can’t show every individual datapoint, it’s impossible. There are however a few things you can do to make me trust what you’re presenting, and they involve making substantial use of Supplementary Figures and Tables. The examples below are going to be proteomics heavy, but I hope you’ll be able to see how you could adapt them relatively easily for other ‘omics techniques.
First off, let’s see some experiment wide distributions. In proteomics, this is usually a sample by sample boxplot of all protein abundances. This way you’ll instantly get an idea of how much normalization you might need for this dataset, and whether any samples just downright failed. How many proteins did you detect across the experiment, and how many were seen in every sample? Let’s see where your missing values are coming from – are they random, or are they related to an important experimental variable? You can try to use a Principal Component Analysis to show that your variable of interest is related to the big sources of variation in the data, but particularly in complex tissues this might not be the case (and that’s okay!). A PCA may highlight some technical variability that you hadn’t previously considered, and can help you tidy up your data. If you’ve done any kind of missing value imputation, batch effect reduction or normalisation, show me a plot that proves you’ve achieved what you wanted to with this processing. Only then can you persuade me that your data is ready for modelling and differential expression analysis. There is nothing I hate more than the first figure presented in a proteomics experiment being the enriched gene ontology terms in the data. It makes me distrust everything that came before.
Once you start modelling, it’s really great to provide a number of supplementary tables along the way. Heatmaps and volcano plots have their place, but they work in addition to a whole bunch of other stuff. Individual level raw and processed data can be hugely useful to people wanting to include your data in a meta-analysis down the line, and you’d be amazed how many people don’t provide it. If you work in human research, just make sure you’re not giving away any information that might help identify an individual. You can get guidance from the tissue resource you’ve worked with on this, but things like binning ages can allow you to publish individual level data without fear of identification. Once you’ve finished your modelling, be sure to include a table with model outcomes for all input proteins/genes, not just the ones that you found to be significant. The pValue distribution of all the model outcomes can help someone judge how well your model worked, and the table can help answer a common question in proteomics; “Was my protein not significant for them, or did they just not detect it?”. Finally, I like to submit all the code and data objects required to run it to a code repository, though again, be sure to cover any anonymization required for human subjects before making these repos public.
As the leader of the group, this is the level of attention to detail that you can, and should be demanding from all your experiments and publications.
I’m not sure I’m allowed to advertise, but if you’re reading this thinking that you don’t understand statistics well enough to demand this level of attention from your team, then I strongly recommend you spend some time with Mine Çetinkaya-Rundel and her Coursera course on Data Analysis with R. She starts at the very beginning and will ease you into thinking in distributions, the differences between assumptions, and the basic skills needed to judge whether a model is satisfactory. In my view, adopting these practices will increase general trust in the work you put out and make people more likely to reference and re-use your hard won data. Surely a win in anyone’s book.

Dr Becky Carlyle
Author
Dr Becky Carlyle is an Alzheimer’s Research UK Senior Research Fellow at University of Oxford, and has previously worked in the USA. Becky writes about her experiences of starting up a research lab and progressing into a more senior research role. Becky’s research uses mass-spectrometry to quantify thousands of proteins in the brains and biofluids of people with dementia. Her lab is working on various projects, including work to compare brain tissue from people with dementia from Alzheimer’s Disease, to tissue from people who have similar levels of Alzheimer’s Disease pathology but no memory problems. Becky is also a mum, she runs, drinks herbal tea’s and reads lots of books.

 Print This Post
Print This Post




